Nextflow 
 ¶
¶
There are several ways to track Nextflow pipeline runs and artifacts in LaminDB.
Using nf-lamin (recommended)¶
The nf-lamin Nextflow plugin automatically tracks transforms, runs, and artifacts without modifying pipeline code. It requires a LaminHub account.
1. Store your Lamin API key as a Nextflow secret:
%%bash
nextflow secrets set LAMIN_API_KEY <your-lamin-api-key>
2. Add the plugin to your nextflow.config:
%%groovy
plugins {
id 'nf-lamin'
}
lamin {
instance = "your-org/your-instance"
api_key = secrets.LAMIN_API_KEY
}
3. Run your pipeline:
%%bash
nextflow run <your-pipeline>
After the run, explore the tracked data in LaminHub or via the Python SDK:
import lamindb as ln
ln.Run.get("your-run-uid")
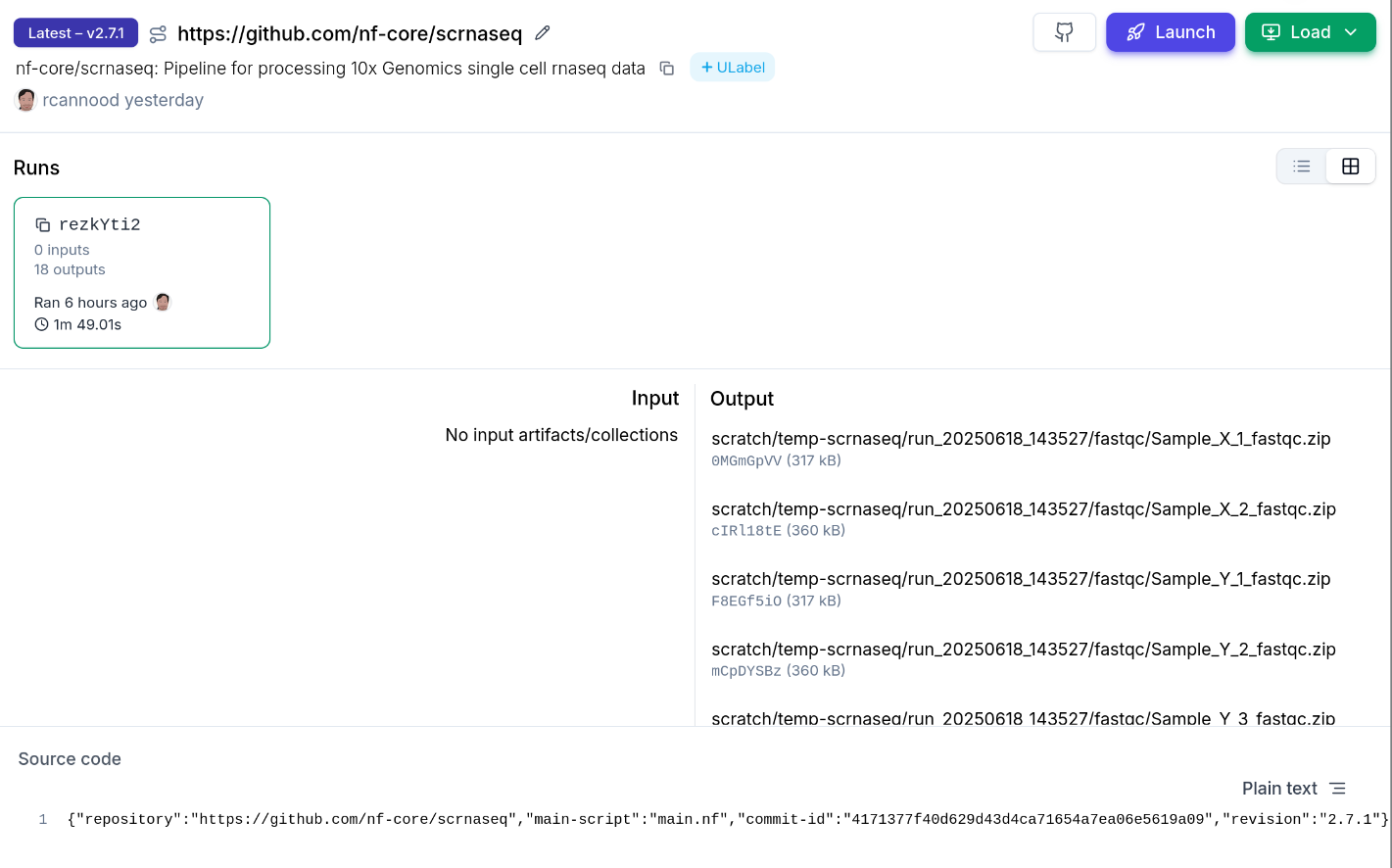
→ See Nextflow: nf-laminfor the fullnf-lamin` configuration reference.
→ See Examples for ready-to-run examples for existing pipelines.
Using a post-run script¶
You can register runs manually without using the nf-lamin plugin using a Python post-run script. First run the pipeline:
# the test profile uses all downloaded input files as an input
!nextflow run nf-core/scrnaseq -r 4.0.0 -profile docker,test -resume --outdir scrnaseq_output
Show code cell output
N E X T F L O W ~ version 25.10.4
Pulling nf-core/scrnaseq ...
downloaded from https://github.com/nf-core/scrnaseq.git
WARN: It appears you have never run this project before -- Option `-resume` is ignored
Downloading plugin nf-schema@2.3.0
Launching `https://github.com/nf-core/scrnaseq` [special_watson] DSL2 - revision: e0ddddbff9 [4.0.0]
------------------------------------------------------
,--./,-.
___ __ __ __ ___ /,-._.--~'
|\ | |__ __ / ` / \ |__) |__ } {
| \| | \__, \__/ | \ |___ \`-._,-`-,
`._,._,'
nf-core/scrnaseq 4.0.0
------------------------------------------------------
Input/output options
input : https://github.com/nf-core/test-datasets/raw/scrnaseq/samplesheet-2-0.csv
outdir : scrnaseq_output
Mandatory arguments
aligner : star
protocol : 10XV2
Skip Tools
skip_cellbender : true
Reference genome options
fasta : https://github.com/nf-core/test-datasets/raw/scrnaseq/nf-lamin/GRCm38.p6.genome.chr19.fa
gtf : https://github.com/nf-core/test-datasets/raw/scrnaseq/nf-lamin/gencode.vM19.annotation.chr19.gtf
save_align_intermeds : true
Institutional config options
config_profile_name : Test profile
config_profile_description: Minimal test dataset to check pipeline function
Generic options
trace_report_suffix : 2026-04-27_13-57-49
Core Nextflow options
revision : 4.0.0
runName : special_watson
containerEngine : docker
launchDir : /home/runner/work/nf-lamin/nf-lamin/docs
workDir : /home/runner/work/nf-lamin/nf-lamin/docs/work
projectDir : /home/runner/.nextflow/assets/nf-core/scrnaseq
userName : runner
profile : docker,test
configFiles : /home/runner/.nextflow/assets/nf-core/scrnaseq/nextflow.config
!! Only displaying parameters that differ from the pipeline defaults !!
------------------------------------------------------
* The pipeline
https://doi.org/10.5281/zenodo.3568187
* The nf-core framework
https://doi.org/10.1038/s41587-020-0439-x
* Software dependencies
https://github.com/nf-core/scrnaseq/blob/master/CITATIONS.md
WARN: The following invalid input values have been detected:
* --monochromeLogs: null
* --validationSchemaIgnoreParams: genomes
[cb/32c855] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:FASTQC_CHECK:FASTQC (Sample_X)
[4c/625f08] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:FASTQC_CHECK:FASTQC (Sample_Y)
[58/494eb7] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:GTF_GENE_FILTER (GRCm38.p6.genome.chr19.fa)
[6a/1dd35e] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:STARSOLO:STAR_GENOMEGENERATE (GRCm38.p6.genome.chr19.fa)
[46/817b2d] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:STARSOLO:STAR_ALIGN (Sample_X)
[b1/176ed0] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:STARSOLO:STAR_ALIGN (Sample_Y)
[8d/246c54] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:MTX_TO_H5AD (Sample_X)
[8a/2a3e08] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:MTX_TO_H5AD (Sample_X)
[9f/5412ce] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:MTX_TO_H5AD (Sample_Y)
[47/fc23d6] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:MTX_TO_H5AD (Sample_Y)
[38/603bda] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:MULTIQC
[3c/a05ac4] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:ANNDATAR_CONVERT (Sample_X)
[6e/209655] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:ANNDATAR_CONVERT (Sample_X)
[dc/976839] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:ANNDATAR_CONVERT (Sample_Y)
[a0/a18b57] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:ANNDATAR_CONVERT (Sample_Y)
[17/e446bc] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:CONCAT_H5AD (combined)
[76/b2ef89] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:CONCAT_H5AD (combined)
[14/cf2f37] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:ANNDATAR_CONVERT (combined)
[f6/1581e7] Submitted process > NFCORE_SCRNASEQ:SCRNASEQ:H5AD_CONVERSION:ANNDATAR_CONVERT (combined)
-[nf-core/scrnaseq] Pipeline completed successfully-
Example: nf-core/scrnaseq
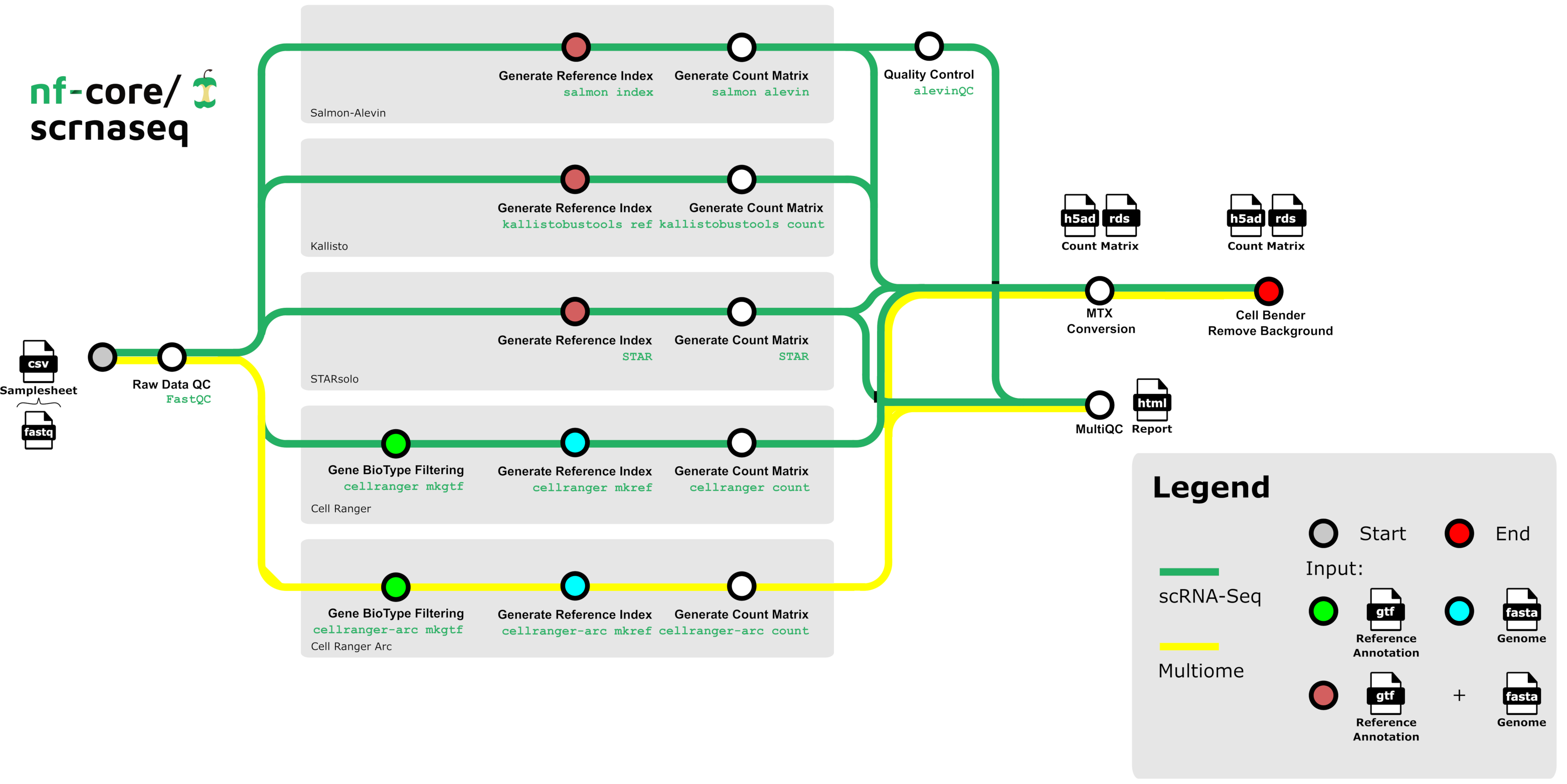
After the run is complete, use a post-run script to register inputs and outputs in LaminDB:
import argparse
import lamindb as ln
import json
import re
from pathlib import Path
from lamin_utils import logger
def parse_arguments() -> argparse.Namespace:
parser = argparse.ArgumentParser()
parser.add_argument("--input", type=str, required=True)
parser.add_argument("--output", type=str, required=True)
return parser.parse_args()
def register_pipeline_io(input_dir: str, output_dir: str, run: ln.Run) -> None:
"""Register input and output artifacts for an `nf-core/scrnaseq` run."""
input_artifacts = ln.Artifact.from_dir(input_dir, run=False)
ln.save(input_artifacts)
run.input_artifacts.set(input_artifacts)
ln.Artifact(f"{output_dir}/multiqc", description="multiqc report", run=run).save()
ln.Artifact(
f"{output_dir}/star/mtx_conversions/combined_filtered_matrix.h5ad",
key="filtered_count_matrix.h5ad",
run=run,
).save()
def register_pipeline_metadata(output_dir: str, run: ln.Run) -> None:
"""Register nf-core run metadata stored in the 'pipeline_info' folder."""
ulabel = ln.ULabel(name="nextflow").save()
run.transform.ulabels.add(ulabel)
# nextflow run id
content = next(Path(f"{output_dir}/pipeline_info").glob("execution_report_*.html")).read_text()
match = re.search(r"run id \[([^\]]+)\]", content)
nextflow_id = match.group(1) if match else ""
run.reference = nextflow_id
run.reference_type = "nextflow_id"
# completed at
completion_match = re.search(r'<span id="workflow_complete">([^<]+)</span>', content)
if completion_match:
from datetime import datetime
timestamp_str = completion_match.group(1).strip()
run.finished_at = datetime.strptime(timestamp_str, "%d-%b-%Y %H:%M:%S")
# execution report and software versions
for file_pattern, description, run_attr in [
("execution_report*", "execution report", "report"),
("nf_core_*_software*", "software versions", "environment"),
]:
matching_files = list(Path(f"{output_dir}/pipeline_info").glob(file_pattern))
if not matching_files:
logger.warning(f"No files matching '{file_pattern}' in pipeline_info")
continue
artifact = ln.Artifact(
matching_files[0],
description=f"nextflow run {description} of {nextflow_id}",
visibility=0,
run=False,
).save()
setattr(run, run_attr, artifact)
# nextflow run parameters
params_path = next(Path(f"{output_dir}/pipeline_info").glob("params*"))
with params_path.open() as params_file:
params = json.load(params_file)
ln.Param(name="params", dtype="dict").save()
run.features.add_values({"params": params})
run.save()
args = parse_arguments()
scrnaseq_transform = ln.Transform(
key="scrna-seq",
version="4.0.0",
type="pipeline",
reference="https://github.com/nf-core/scrnaseq",
).save()
run = ln.Run(transform=scrnaseq_transform).save()
register_pipeline_io(args.input, args.output, run)
register_pipeline_metadata(args.output, run)
!python nextflow/register_scrnaseq_run.py --input scrnaseq_input --output scrnaseq_output
Show code cell output
/home/runner/work/nf-lamin/nf-lamin/docs/nextflow/register_scrnaseq_run.py:77: DeprecationWarning: `type` argument of transform was renamed to `kind` and will be removed in a future release.
scrnaseq_transform = ln.Transform(
Traceback (most recent call last):
File "/home/runner/work/nf-lamin/nf-lamin/docs/nextflow/register_scrnaseq_run.py", line 77, in <module>
scrnaseq_transform = ln.Transform(
~~~~~~~~~~~~^
key="scrna-seq",
^^^^^^^^^^^^^^^^
...<2 lines>...
reference="https://github.com/nf-core/scrnaseq",
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
).save()
^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/lamindb/models/transform.py", line 305, in __init__
.first()
~~~~~^^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/lamindb/models/query_set.py", line 1309, in first
if len(self) == 0:
~~~^^^^^^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/django/db/models/query.py", line 368, in __len__
self._fetch_all()
~~~~~~~~~~~~~~~^^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/django/db/models/query.py", line 1954, in _fetch_all
self._result_cache = list(self._iterable_class(self))
~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/django/db/models/query.py", line 93, in __iter__
results = compiler.execute_sql(
chunked_fetch=self.chunked_fetch, chunk_size=self.chunk_size
)
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/django/db/models/sql/compiler.py", line 1623, in execute_sql
cursor.execute(sql, params)
~~~~~~~~~~~~~~^^^^^^^^^^^^^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/django/db/backends/utils.py", line 79, in execute
return self._execute_with_wrappers(
~~~~~~~~~~~~~~~~~~~~~~~~~~~^
sql, params, many=False, executor=self._execute
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
)
^
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/django/db/backends/utils.py", line 92, in _execute_with_wrappers
return executor(sql, params, many, context)
File "/opt/hostedtoolcache/Python/3.14.4/x64/lib/python3.14/site-packages/lamindb_setup/core/django.py", line 39, in error_no_instance_wrapper
raise CurrentInstanceNotConfigured
lamindb_setup.errors.CurrentInstanceNotConfigured: No instance is connected! Call
- CLI: lamin connect / lamin init
- Python: ln.connect() / ln.setup.init()
- R: ln$connect() / ln$setup$init()
Such a script can also be triggered from a serverless environment (e.g., AWS Lambda).